Under the Microscope
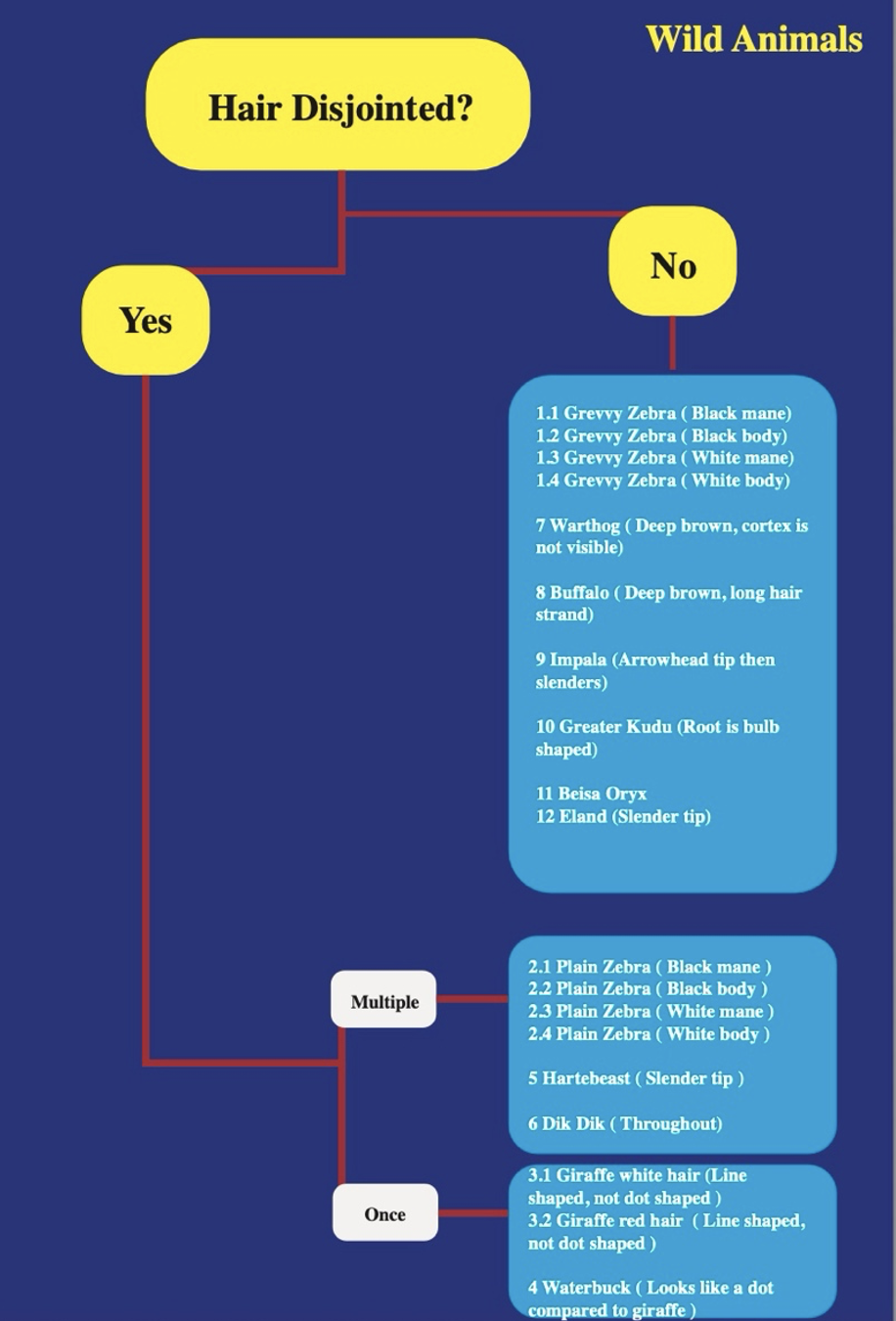
Hair structure as a species identifier
Each animal species has a uniquely structured hair strand. Differentiation requires careful analysis of the medulla (the innermost, darker section) and the cortex (the outer layer) — looking at size, colour, breakage patterns, and overall morphology.
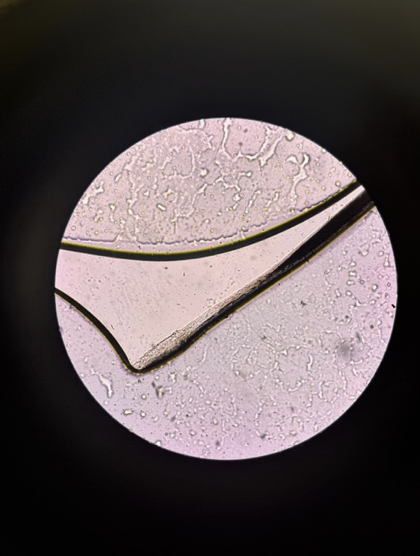
Single hair strand under microscope — the dark inner section is the medulla, surrounded by the lighter cortex layer.
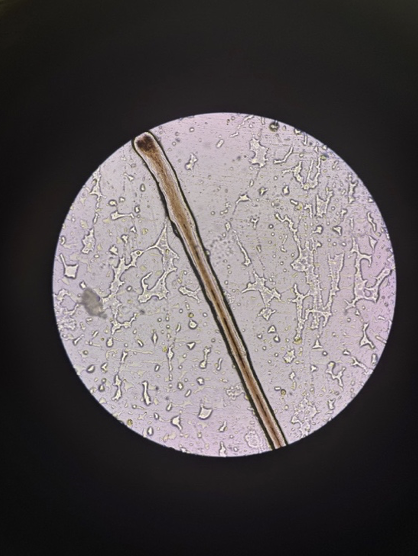
Multiple strands mounted together — comparing structure, thickness, and tip morphology across specimens from the same sample.